Abstract
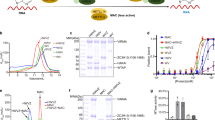
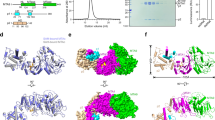
N6-methyladenosine (m6A) is the most abundant internal modification of nearly all eukaryotic mRNAs and has recently been reported to be recognized by the YTH domain family proteins. Here we present the crystal structures of the YTH domain of YTHDC1, a member of the YTH domain family, and its complex with an m6A-containing RNA. Our structural studies, together with transcriptome-wide identification of YTHDC1-binding sites and biochemical experiments, not only reveal the specific mode of m6A-YTH binding but also explain the preferential recognition of the GG(m6A)C sequences by YTHDC1.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
19 August 2015
In the version of this Brief Communication initially published, the email address for corresponding author, Chao Xu, was incorrect. It should be listed as chaor.xu@utoronto.edu. This error has been corrected in the PDF and HTML versions of the article.
11 November 2015
In the version of this Brief Communication initially published, as well as in a corrected version of 19 August 2015, incorrect e-mail addresses were given for corresponding author Chao Xu. The correct address is chaor.xu@utoronto.ca. This error has been corrected in the PDF and HTML versions of the article.
References
Tuck, M.T. Int. J. Biochem. 24, 379–386 (1992).
Schwartz, S. et al. Cell 155, 1409–1421 (2013).
Wei, C.M. & Moss, B. Biochemistry 16, 1672–1676 (1977).
Zhong, S. et al. Plant Cell 20, 1278–1288 (2008).
Bokar, J.A. Fine-tuning of RNA functions by modification and editing. Vol. 12 (ed. Grosjean, H.) 141–177 (Springer-Verlag, Berlin Heidelberg, 2005).
Hongay, C.F. & Orr-Weaver, T.L. Proc. Natl. Acad. Sci. USA 108, 14855–14860 (2011).
Dominissini, D. et al. Nature 485, 201–206 (2012).
Meyer, K.D. et al. Cell 149, 1635–1646 (2012).
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Nat. Rev. Genet. 15, 293–306 (2014).
Jia, G. et al. Nat. Chem. Biol. 7, 885–887 (2011).
Zheng, G. et al. Mol. Cell 49, 18–29 (2013).
Liu, J. et al. Nat. Chem. Biol. 10, 93–95 (2014).
Wang, Y. et al. Nat. Cell Biol. 16, 191–198 (2014).
Wang, X. et al. Nature 505, 117–120 (2014).
Zhang, Z. et al. J. Biol. Chem. 285, 14701–14710 (2010).
Nayler, O., Hartmann, A.M. & Stamm, S. J. Cell Biol. 150, 949–962 (2000).
Zhang, B. et al. Int. J. Gynecol. Cancer 20, 492–499 (2010).
Hirschfeld, M. et al. Mol. Carcinog. 10.1002/mc.22045 (13 June 2013).
Hafner, M. et al. Cell 141, 129–141 (2010).
Jacobs, S.A. & Khorasanizadeh, S. Science 295, 2080–2083 (2002).
Wang, G.G. et al. Nature 459, 847–851 (2009).
Zou, X., Ma, W., Solov'yov, I.A., Chipot, C. & Schulten, K. Nucleic Acids Res. 40, 2747–2758 (2012).
Marcotrigiano, J., Gingras, A.C., Sonenberg, N. & Burley, S.K. Cell 89, 951–961 (1997).
Kabsch, W. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P.R. & Murshudov, G.N. Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
McCoy, A.J. et al. J. Appl. Crystallogr. 40, 658–674 (2007).
Lebedev, A.A. et al. Acta Crystallogr. D Biol. Crystallogr. 68, 431–440 (2012).
Murshudov, G.N. et al. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Chen, V.B. et al. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Hafner, M. et al. J. Vis. Exp. 2 10.3791/2034 (2010).
Pearson, W.R., Wood, T., Zhang, Z. & Miller, W. Genomics 46, 24–36 (1997).
Trapnell, C., Pachter, L. & Salzberg, S.L. Bioinformatics 25, 1105–1111 (2009).
Corcoran, D.L. et al. Genome Biol. 12, R79 (2011).
Acknowledgements
We would like to thank J. Walker for assistance in structure determination. The SGC is a registered charity (number 1097737) that receives funds from AbbVie, Boehringer Ingelheim, the Canada Foundation for Innovation, the Canadian Institutes for Health Research, Genome Canada through the Ontario Genomics Institute (OGI-055), GlaxoSmithKline, Janssen, Lilly Canada, the Novartis Research Foundation, the Ontario Ministry of Economic Development and Innovation, Pfizer, Takeda and the Wellcome Trust (092809/Z/10/Z to J.M.). C.H. is supported by the Howard Hughes Medical Institute as an investigator. The Mass Spectrometry Facility of the University of Chicago is funded by the National Science Foundation (CHE-1048528).
Author information
Authors and Affiliations
Contributions
C.X. and J.M. conceived the project; C.X. performed the structural and binding experiments with assistance from K.L., W.T. and Y.L.; X.W. and I.A.R. conducted the PAR-CLIP experiment of YTHDC1; Z.L. and X.W. analyzed the PAR-CLIP data. C.X., X.W., C.H. and J.M. wrote the manuscript. All authors contributed to editing the manuscript. C.H. and J.M. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1 and 2 and Supplementary Figures 1–12. (PDF 2897 kb)
Rights and permissions
About this article
Cite this article
Xu, C., Wang, X., Liu, K. et al. Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain. Nat Chem Biol 10, 927–929 (2014). https://doi.org/10.1038/nchembio.1654
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1654
This article is cited by
-
The role of the methyltransferase METTL3 in prostate cancer: a potential therapeutic target
BMC Cancer (2024)
-
ChemRAP uncovers specific mRNA translation regulation via RNA 5′ phospho-methylation
EMBO Reports (2024)
-
The PIN1-YTHDF1 axis promotes breast tumorigenesis via the m6A-dependent stabilization of AURKA mRNA
Archives of Pharmacal Research (2024)
-
Small RNA modifications: regulatory molecules and potential applications
Journal of Hematology & Oncology (2023)
-
O-GlcNAcylation determines the translational regulation and phase separation of YTHDF proteins
Nature Cell Biology (2023)