Abstract
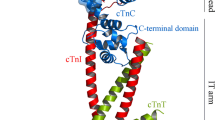
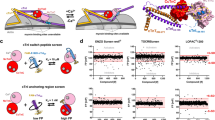
The Ca2+-sensitive cardiac troponin (cTn) is a hetero-trimer complex consisting of three subunits cTnC, cTnI, and cTnT, which has been recognized as an important biomarker and a potential target of cardiovascular diseases. Previously, several small-molecule agents such as levosimendan and pimobendan have been successfully developed to target this protein for the treatment of heart failure. Here, instead of small-molecule chemical drugs, we purposed rational derivation of self-inhibitory peptides as potential biologic disruptors of cTnC–cTnI interaction from the interaction complex interface. In the procedure, the crystal structure of cTn trimer was examined in detail using bioinformatics approach, from which a peptide-mediated interaction between the N-terminal domain of cTnC and the switch region of cTnI was identified. The switch is a 19-mer peptide segment Swt that contains a structured helical core capped by a short N-terminal tripeptide and a disordered C-terminal tail. Structural and energetic analysis revealed that the Swt peptide binds independently to cTnC N-terminal domain, which can be stripped from the intact cTnI subunit to interact effectively with cTnC. Further investigations found that truncation of two N-terminal residues and five C-terminal residues of the full-length Swt peptide, resulting in a shortened version namely Swt-ΔN2ΔC5 peptide, would not cause substantial loss in its binding potency to cTnC. The computational finding was then confirmed by using fluorescence-based affinity assays; the Swt and Swt-ΔN2ΔC5 peptides was experimentally measured to have a moderately high affinity to the recombinant protein of human cTnC N-terminal domain with K d values at micromolar level. The Swt and Swt-ΔN2ΔC5 are considered as inhibitory peptides that can be further optimized and modified to obtain high-affinity disruptors of cTnI–cTnC interaction.



Similar content being viewed by others
References
Bahar I, Lezon TR, Bakan A, Shrivastava IH (2010) Normal mode analysis of biomolecular structures: functional mechanisms of membrane proteins. Chem Rev 110:1463–1497
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Sh IN, dyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 2:235–242
Chen D, Liu S, Zhang W, Sun L (2015) Rational design of YAP WW1 domain-binding peptides to target TGFβ/BMP/Smad-YAP interaction in heterotopic ossification. J Pept Sci 21:826–832
Darden T, York D, Pedersen L (1983) Particale mesh Ewald and N.log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Davis JP, Tikunova SB (2008) Ca2+ exchange with troponin C and cardiac muscle dynamics. Cardiovasc Res 77:619–626
Fitton A, Brogden RN (1994) Pimobendan. a review of its pharmacology and therapeutic potential in congestive heart failure. Drugs Aging 4:417–441
Gordon JC, Myers JB, Folta T, Shoja V, Heath LS, Onufriev A (2005) H++: a server for estimating pKa’s and adding missing hydrogens to macromolecules. Nucleic Acids Res 33:W368–W371
Haikala H, Kaivola J, Nissinen E, Wall P, Levijoki J, Lindén IB (1995) Cardiac troponin C as a target protein for a novel calcium sensitizing drug, levosimendan. J Mol Cell Cardiol 27:1859–1866
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: A linear constraint solver for molecular simulations. J Comput Chem 18:1463–1472
Homeyer N, Gohlke H (2012) Free energy calculations by the molecular mechanics Poisson-Boltzmann surface area method. Mol Inf 31:114–122
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Phys Chem 79:926–935
Ko J, Park H, Heo L, Seok C (2012) GalaxyWEB server for protein structure prediction and refinement. Nucleic Acids Res 40:W294–W297
Li MX, Wang X, Sykes BD (2004) Structural based insights into the role of troponin in cardiac muscle pathophysiology. J Muscle Res Cell Motil 25:559–579
London N, Raveh B, Movshovitz-Attias D, Schueler-Furman O (2010) Can self-inhibitory peptides be derived from the interfaces of globular protein-protein interactions? Proteins 78:3140–3149
Potluri S, Ventura HO, Mulumudi M, Mehra MR (2004) Cardiac troponin levels in heart failure. Cardiol Rev 12:21–25
Sedan Y, Marcu O, Lyskov S, Schueler-Furman O (2016) Peptiderive server: derive peptide inhibitors from protein-protein interactions. Nucleic Acids Res 44:W536–W541
Sharma S, Jackson PG, Makan J (2004) Cardiac troponins. J Clin Pathol 57:1025–1026
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
White SP, Cohen C, Phillips GN (2002) Structure of co-crystals of tropomyosin and troponin. Nature 325:826–828
Word JM, Lovell SC, Richardson JS, Richardson DC (1999) Asparagine and glutamine: using hydrogen atom contacts in the choice of side-chain amide orientation. J Mol Biol 285:1735–1747
Yang C, Zhang S, He P, Wang C, Huang J, Zhou P (2015) Self-binding peptides: folding or binding? J Chem Inf Model 55:329–342
Yu H, Zhou P, Deng M, Shang Z (2014) Indirect readout in protein-peptide recognition: a different story from classical biomolecular recognition. J Chem Inf Model 54:2022–2032
Acknowledgements
This work was supported by the Shanghai Chest Hospital.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Human and Animal Rights
Not applicable.
Informed Consent
Not applicable.
Rights and permissions
About this article
Cite this article
Xu, Y., Huang, R., Gu, J. et al. Derivation of Inhibitory Peptides to Target the Cardiac Troponin C–I Interaction as Potential Therapeutics for Heart Failure. Int J Pept Res Ther 23, 387–392 (2017). https://doi.org/10.1007/s10989-017-9576-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-017-9576-6